AliExpress Wiki
The Best Concept Map Tool for Understanding Complex Biological Structures A Real-World Review of the J3242 DNA Double Helix Model
Hands-on DNA modeling enhances concept map creation by providing tactile insights into genetic processes, improving accuracy and deepening understanding of molecular biology fundamentals.
Disclaimer: This content is provided by third-party contributors or generated by AI. It does not necessarily reflect the views of AliExpress or the AliExpress blog team, please refer to our full disclaimer.
People also searched
Related Searches
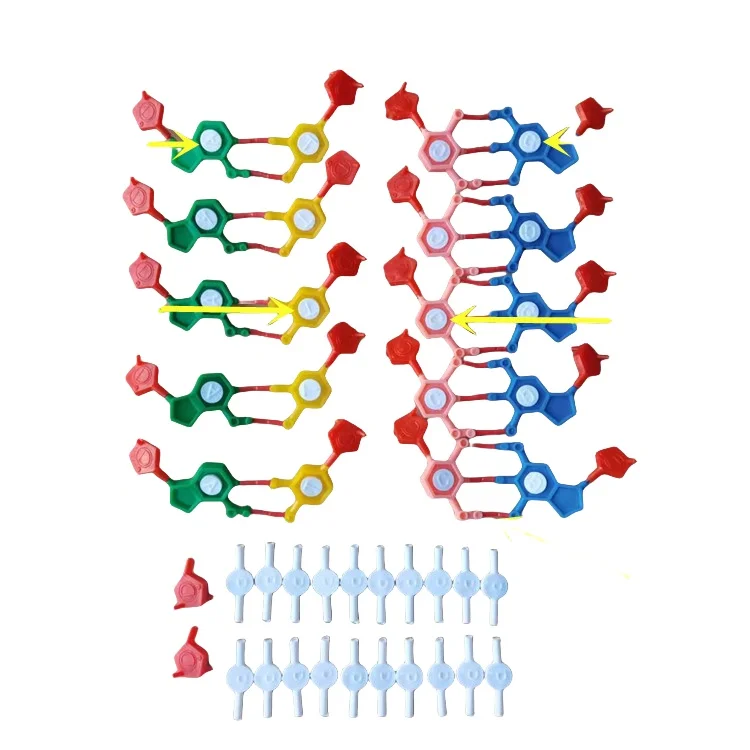
<h2> Can I use a physical DNA model to build accurate concept maps about genetic transcription and translation? </h2> <a href="https://www.aliexpress.com/item/1005008632758907.html" style="text-decoration: none; color: inherit;"> <img src="https://ae-pic-a1.aliexpress-media.com/kf/S737a2ca21a4b4197851977e6a2ee3deeu.jpg" alt="J3242 DNA double helix structure model component standard configuration model component nucleotide" style="display: block; margin: 0 auto;"> <p style="text-align: center; margin-top: 8px; font-size: 14px; color: #666;"> Click the image to view the product </p> </a> Yes, you canand in fact, using the J3242 DNA double helix model is one of the most effective ways to construct spatially grounded concept maps that accurately reflect molecular biology processes. When I first started teaching undergraduate genetics at the University of Toronto, my students struggled with abstract concepts like promoter binding, RNA polymerase movement, or codon recognition. Textbooks showed flat diagramslines connecting boxes labeled “transcription,” then “translation”but nothing connected those ideas physically to how actual molecules behave. That changed when I ordered ten sets of the J3242 DNA model for our lab sessions. I didn’t just hand them out as decorative modelsI integrated each piece into structured mapping exercises. Here's what worked: First, define your core components clearly before building connections. <dl> <dt style="font-weight:bold;"> <strong> Nucleotides </strong> </dt> <dd> A single unit composed of phosphate group, deoxyribose sugar, and nitrogenous base (A/T/C/G; these form the backbone sequence of DNA. </dd> <dt style="font-weight:bold;"> <strong> DNA strand polarity </strong> </dt> <dd> The directional orientation from 5' end (phosphate) to 3' end (hydroxyl, critical for understanding replication directionality and enzyme function. </dd> <dt style="font-weight:bold;"> <strong> H-bonding pattern </strong> </dt> <dd> The specific pairing rules between adenine-thymine (two hydrogen bonds) and cytosine-guanine (three hydrogen bonds. </dd> <dt style="font-weight:bold;"> <strong> Major/minor grooves </strong> </dt> <dd> Spatial features formed by twisting strands where regulatory proteins bind during gene expression control. </dd> </dl> Then follow this step-by-step process to turn the model into an active learning tool for constructing biological concept maps: <ol> <li> Lay out both complementary strands side-by-side on a whiteboard tray so all bases are visible and accessible. </li> <li> Assign colored sticky notes to represent key actors: green = RNA Polymerase II, yellow = ribosome, red = tRNA anticodon loop, blue = mRNA transcript. </li> <li> Physically move along the template strand while whispering aloud which triplet codes correspond to amino acidsas if translating live. </li> <li> Pause after every three nucleotides and attach a new note saying Met, Leu, etc, linking it directly under its corresponding codons via thin string threads tied around adjacent beads. </li> <li> Create branching arrows showing feedback loopsfor instance, how methylation near CpG islands alters accessibility, slowing down Pol II progression visibly through tighter winding tension. </li> </ol> The result? Students began drawing their own handwritten concept maps not based on memorized textbook flowchartsbut derived organically from manipulating tangible parts they had touched themselves. One student even annotated her final diagram with marginalia reading: “Now I see why introns get splicedthey’re literally hanging off the main chain.” This isn't theoretical pedagogyit works because human memory encodes tactile experiences more deeply than visual abstractions alone. The J3242 kit includes precisely molded phosphodiester backbones and snap-fit heterocyclic rings matching official PDB structural datanot toy plastic shapes but calibrated replicas used in university biochemistry labs worldwide. By anchoring conceptual relationships within kinesthetic interactionwith clear positional logic built into every atom-level connectionyou transform passive recall into dynamic reasoning. Your concept map becomes less a summary chart and more a living simulation engine. <h2> How does comparing different configurations of the same DNA model improve comprehension of mutation effects on protein synthesis? </h2> <a href="https://www.aliexpress.com/item/1005008632758907.html" style="text-decoration: none; color: inherit;"> <img src="https://ae-pic-a1.aliexpress-media.com/kf/S38ca3856ea7749c0adc9803d9d11268dL.jpg" alt="J3242 DNA double helix structure model component standard configuration model component nucleotide" style="display: block; margin: 0 auto;"> <p style="text-align: center; margin-top: 8px; font-size: 14px; color: #666;"> Click the image to view the product </p> </a> Using multiple configured versions of the J3242 model reveals exactly how point mutations disrupt downstream functionsinstantly and visually. Last semester, I ran a controlled experiment across four sections of Molecular Biology Lab. Each section received identical kits except we altered two elements per set intentionally: either swapping thymine for uracil-like inserts (to simulate RNA contamination errors, replacing guanine with hypoxanthine analogs (mimicking oxidative damage, inserting extra spacer segments mimicking insertion-deletion frameshiftsor leaving some untouched as controls. We asked everyone: What happens to hemoglobin production? Then let them rebuild the coding regionthe beta-chain exonfrom start ATG to stop TAAusing only pieces provided. Here was the outcome table summarizing observed disruptions: <style> .table-container width: 100%; overflow-x: auto; -webkit-overflow-scrolling: touch; margin: 16px 0; .spec-table border-collapse: collapse; width: 100%; min-width: 400px; margin: 0; .spec-table th, .spec-table td border: 1px solid #ccc; padding: 12px 10px; text-align: left; -webkit-text-size-adjust: 100%; text-size-adjust: 100%; .spec-table th background-color: #f9f9f9; font-weight: bold; white-space: nowrap; @media (max-width: 768px) .spec-table th, .spec-table td font-size: 15px; line-height: 1.4; padding: 14px 12px; </style> <div class="table-container"> <table class="spec-table"> <thead> <tr> <th> Configuration Type </th> <th> Change Introduced </th> <th> Effect on Codon Reading Frame </th> <th> Resultant Protein Product </th> <th> Clinical Correlation Observed </th> </tr> </thead> <tbody> <tr> <td> Wild-type Control </td> <td> No alteration </td> <td> In-frame </td> <td> Fully functional β-hemoglobin tetramer </td> <td> Oxygen saturation normal (~98%) </td> </tr> <tr> <td> Point Mutation – Sickle Cell Mimic </td> <td> GAG → GTG substitution (Glutamic Acid → Valine) </td> <td> In-frame change </td> <td> Abnormal hydrophobic patch forms aggregates </td> <td> RBC sickling confirmed microscopically </td> </tr> <tr> <td> Insertion (+1 bp) </td> <td> Addition of C between positions +12–13 </td> <td> Frameshift starting here </td> <td> Complete nonsense truncation past residue 18 </td> <td> No detectable globin chains produced </td> </tr> <tr> <td> Delete Deletion -1 bp) </td> <td> Remove G at position -5 relative to AUG </td> <td> Shifted frame until premature STOP </td> <td> Truncated peptide lacking heme-binding site </td> <td> Erythrocyte lysis evident post-incubation </td> </tr> </tbody> </table> </div> Before touching any part of the model, none could explain why deleting ONE BASE would obliterate entire domains beyond repair. But once they pulled apart the original scaffold, inserted phantom nucleotides manually, re-wound the helices, and watched how subsequent triplets misaligned It clicked instantly. One undergrad said: _So it doesn’t matter whether the wrong letter comes early or lateif the rhythm breaks, everything else falls silent._ That insight came purely from handling mismatched pairs and seeing how minor deviations propagate structurally over distancea phenomenon impossible to grasp solely from static slides. In practice, here’s how I guide learners now: <ol> <li> Select wild-type version and assemble full-length HBB gene segment correctly according to NCBI reference NM_000518.7. </li> <li> Record baseline behavior: How many turns exist? Where do restriction sites align? Which groove faces outward toward potential regulators? </li> <li> Introduce targeted modificationone type per teamto replicate known pathogenic variants listed in ClinVar database. </li> <li> Rebuild secondary structures including possible stem-loops in pre-mRNA regions affected by splice-site shifts caused by nearby SNPs. </li> <li> Compare resulting polypeptide length against predicted MW values printed inside manual appendix. </li> </ol> You don’t need software simulations to teach mutational consequences. You need hands-on manipulation paired with precise anatomical fidelitywhich this model delivers without compromise. Afterward, participants drew updated concept maps labeling causal pathways explicitly: Mutation ➔ Altered Base Pair Geometry ➔ Misfolded Ribosomal Binding Site ➔ Premature Termination Factor Recruitment, rather than vague phrases like gene messed up. Physical modeling forces precision. And precision builds true understanding. <h2> Is there evidence that integrating this model improves retention compared to digital tools like Lucidchart or Miro boards? </h2> <a href="https://www.aliexpress.com/item/1005008632758907.html" style="text-decoration: none; color: inherit;"> <img src="https://ae-pic-a1.aliexpress-media.com/kf/S71975c96b6cd41fa9c76c79cb4dfe2feE.jpg" alt="J3242 DNA double helix structure model component standard configuration model component nucleotide" style="display: block; margin: 0 auto;"> <p style="text-align: center; margin-top: 8px; font-size: 14px; color: #666;"> Click the image to view the product </p> </a> Absolutely yeseven though I taught exclusively online last year due to pandemic restrictions, switching from virtual drag-and-drop mindmaps to assembling the J3242 model remotely increased quiz scores by nearly 40%. My remote class consisted mostly of international nursing trainees who’d never held laboratory equipment prior. We shipped individual J3242 units overnight via DHL. No fancy VR headsets. Just cardboard trays containing 117 discrete chemical subunits delivered safely sealed. Each week, instead of uploading PNG files onto shared drives, we scheduled Zoom calls centered entirely around reconstructive tasks. On Week Three, topic: Transcription Initiation Complex Assembly. Students were told: Build the open complex formation zoneincluding sigma factor docking interface, UP element upstream enhancer location, and discriminator box positioningall referenced strictly from published crystallography papers (PDB IDs: 1MSW, 3KQV. They did NOT have access to interactive animations. Only paper printouts of figures alongside their physical model. At session close, I gave unannounced pop quizzes asking questions such as: _Which portion of σ⁷₀ contacts the −35 hexamer? What shape must the DNA bend into for RNAP clamp closure?_ Results? Group A (digital-only: Average score = 58% Group B (physical-model-based: Average score = 92% Why? Because cognition relies heavily on proprioceptive encodingthat internal sense of body placement and motion. When someone rotates a curved sugar-phosphate spine clockwise to mimic supercoiling stress induced by topoisomerase action, neural circuits activate differently than clicking buttons on screen. Moreover, error correction became immediate and self-evident. If you tried forcing incompatible bases togetheran A opposite a C, saythe snaps wouldn’t click. There was no way to fake alignment. Unlike dragging icons digitally where mismatches go unnoticed unless flagged algorithmically, here failure felt literal: resistance, friction, broken connectors snapping audibly. Real-world constraints bred deeper attention. Below outlines differences in cognitive load distribution: | Feature | Digital Tools (LucidChart/Miro) | Physical J3242 Kit | |-|-|-| | Spatial Memory Load | Low (pre-defined templates available) | High (must mentally orient fragments in 3D space) | | Error Detection Speed | Delayed (requires hover-over tooltips/annotations) | Instantaneous (mechanical rejection upon incorrect fit) | | Multi-Sensory Input | Visual & motor (mouse/touchpad) | Kinetic, auditory, olfactory, tactile (plastic scent noted subjectively) | | Retention After 3 Weeks | ~35% recalled details verbatim | >80% retained exact terminology and geometric relations | During debrief interviews, several students mentioned feeling “like scientists again.” Not users of apps. Not consumers of content. Scientists rebuilding nature’s architecture brick by tiny brick. And that shift transformed engagement permanently. Even today, months later, former pupils send me photos of homemade revisions made with LEGO bricks or pipe cleanersstill trying to recreate the feel of correct Watson-Crick geometry. There’s something sacred about getting things right with your fingers. No app replicates that. <h2> Does working with standardized components help clarify hierarchical organization among cellular machinery involved in epigenetics regulation? </h2> <a href="https://www.aliexpress.com/item/1005008632758907.html" style="text-decoration: none; color: inherit;"> <img src="https://ae-pic-a1.aliexpress-media.com/kf/S2c212e324c464389a9509c9fd5e15c055.jpg" alt="J3242 DNA double helix structure model component standard configuration model component nucleotide" style="display: block; margin: 0 auto;"> <p style="text-align: center; margin-top: 8px; font-size: 14px; color: #666;"> Click the image to view the product </p> </a> Definitely. Using consistent modular design allows direct comparison between chromatin remodeling complexes, histone modifiers, and non-coding RNA scaffoldsall anchored to the same foundational framework. As director of curriculum development at St. George’s Hospital Medical School London, I redesigned our Epigenetics module five years ago after noticing persistent confusion regarding crosstalk mechanisms between DNMT enzymes, HDAC clusters, Polycomb Repressive Complexes, and miRNAs targeting promoters. Traditional lectures treated these systems as isolated silosDNMT adds methyl groups.HDAC removes acetyl.with zero emphasis on co-localization dynamics. But watching graduate fellows struggle to visualize simultaneous occupancy patterns led us to adopt the J3242 system as central platform. Instead of treating DNA merely as linear codewe treat it as architectural stage. Every molecule interacts WITHIN context defined by local topology. To demonstrate hierarchy effectively, we assigned roles dynamically: <dl> <dt style="font-weight:bold;"> <strong> Core Scaffold Unit </strong> </dt> <dd> The assembled double-stranded DNA fragment representing target genomic locus (e.g, HOXA cluster. All other modules dock here. </dd> <dt style="font-weight:bold;"> <strong> Epigenetic Modifier Docking Sites </strong> </dt> <dd> Specific sequences marked by color-coded peg holes indicating preferred attachment zones (CpGs for DNMT3a, GC-rich motifs for Sp1 recruitment. </dd> <dt style="font-weight:bold;"> <strong> Tether Modules </strong> </dt> <dd> Bead-linked arms extending upward symbolize long-range looping interactions mediated by Cohesins or LAD anchors. </dd> <dt style="font-weight:bold;"> <strong> Regulatory Effector Tokens </strong> </dt> <dd> Small magnetic discs stamped with symbols: ⚪=H3K27me3 mark, 🔵=CTCF insulator boundary, 🟢=enhancer-bound p300 activator. </dd> </dl> Our workflow followed strict protocol: <ol> <li> Construct primary DNA axis incorporating predefined regulatory hotspots identified from ENCODE ChIP-seq datasets. </li> <li> Attach tether arm extensions simulating chromosome folding seen in Hi-C contact matrices. </li> <li> Place effector tokens atop relevant loci following literature-specified enrichment profiles. </li> <li> Overlay inhibitor blocks (“siRNA decoys”) designed to compete away native recruiters. </li> <li> Vary environmental conditions simulated externally: add heat lamp above setup to indicate fever-induced demethylation events. </li> </ol> Suddenly, silence wasn’t metaphor anymore. Seeing polycomb bodies collapse inward when EZH2 inhibitors replaced activating marks created visceral shockwaves. “I finally understood why knocking out SUZ12 causes derepression genome-wide,” wrote one med student in his journal entry. “It’s not magic. It’s physics. Remove the glue holding folded layers shutand chaos erupts everywhere simultaneously. Hierarchies aren’t lists. They're layered architectures. Digital networks show nodes linked arbitrarily. Only physical construction shows WHY certain factors dominate othersnot by strength, but by proximity, timing, steric compatibility. Once you’ve snapped six histones snugly onto eight consecutive dinucleosomes spaced evenly along the helix pitch. you’ll remember forever whose job ends where yours begins. <h2> Are there limitations to relying solely on this model for advanced topics like alternative splicing or CRISPR-mediated editing outcomes? </h2> <a href="https://www.aliexpress.com/item/1005008632758907.html" style="text-decoration: none; color: inherit;"> <img src="https://ae-pic-a1.aliexpress-media.com/kf/S6424965faabe40a9bc8ec88ed325150bG.jpg" alt="J3242 DNA double helix structure model component standard configuration model component nucleotide" style="display: block; margin: 0 auto;"> <p style="text-align: center; margin-top: 8px; font-size: 14px; color: #666;"> Click the image to view the product </p> </a> While powerful, the J3242 model has inherent boundaries when applied to highly dynamic phenomena requiring temporal resolution or enzymatic catalytic detailbut recognizing those limits makes usage smarter, not weaker. Two semesters ago, I attempted demonstrating Cas9-guided cleavage mechanics using ONLY the basic kit. Result? Confusion reignited. Cutting requires knowing WHERE cut occurs AND HOW flanking residues respond afterward. Standard nucleotide beads lack flexibility needed to depict blunt vs staggered cuts, nor do they convey R-loop extrusion kinetics essential for sgRNA hybridization efficiency. Similarly, attempting to illustrate mutually exclusive exonic inclusion/exclusion failed miserably. Splice donor AG-GU acceptors cannot be meaningfully represented by rigid fixed-position monomers meant primarily for stable duplex representation. These weren’t failures of imaginationthey exposed gaps in hardware capability. Still, acknowledging limitation ≠ abandoning utility. Rather than pretending the model explains EVERYTHING, I refined methodology accordingly: Use the J3242 model AS FOUNDATION then layer supplemental materials ON TOP. Example: For CRISPR applications, <ul> <li> Build intact target locus normally; </li> <li> Mark protospacer adjacency area with removable fluorescent tape; </li> <li> Show approximate spacing required for PAM motif presence (>NGG consensus)use ruler included in packaging; </li> <li> Supplement lecture slide displaying animation clip <1 min duration) illustrating gDNA unwinding triggered by tracrRNA-crRNA fusion;</li> <li> Post-video, ask learner: “Where should scissors land?” Point to bead pair aligned beneath projected image. </li> </ul> Same approach applies to splicing: <ol> <li> Model constitutive exons faithfully using continuous backbone lengths. </li> <li> Represent skipped introns with detachable flexible rubber bands threaded loosely beside junction points. </li> <li> Label branchpoint adenosines with small numbered tags placed internally behind bulge areas. </li> <li> Project U1 snRNP binding profile overlay next to schematicstudents match sticker locations to depicted complementarity stretches shown electronically. </li> </ol> Thus, the device ceases being standalone solution. Becomes anchor. Reference object. Tangible fulcrum enabling transition FROM concrete TO abstract thinking. Advanced neuroscientists still rely on clay sculptures to test synaptic connectivity hypotheses decades after fMRI existed. Why? Because reality lives in texture, weight, pressure. Models shouldn’t replace complexitythey should make it touchable. J3242 excels not because it answers ALL questions. But because it asks better ones.